cat_plot is a complementary function to interact_plot() that is designed
for plotting interactions when both predictor and moderator(s) are
categorical (or, in R terms, factors).
Usage
cat_plot(
model,
pred,
modx = NULL,
mod2 = NULL,
data = NULL,
geom = c("point", "line", "bar"),
pred.values = NULL,
modx.values = NULL,
mod2.values = NULL,
interval = TRUE,
plot.points = FALSE,
point.shape = FALSE,
vary.lty = FALSE,
centered = "all",
int.type = c("confidence", "prediction"),
int.width = 0.95,
line.thickness = 1.1,
point.size = 1.5,
pred.point.size = 3.5,
jitter = 0.1,
geom.alpha = NULL,
dodge.width = NULL,
errorbar.width = NULL,
interval.geom = c("errorbar", "linerange"),
outcome.scale = "response",
robust = FALSE,
cluster = NULL,
vcov = NULL,
pred.labels = NULL,
modx.labels = NULL,
mod2.labels = NULL,
set.offset = 1,
x.label = NULL,
y.label = NULL,
main.title = NULL,
legend.main = NULL,
colors = NULL,
partial.residuals = FALSE,
point.alpha = 0.6,
color.class = NULL,
at = NULL,
...
)Arguments
- model
A regression model. The function is tested with
lm,glm,svyglm,merMod,rq,brmsfit,stanregmodels. Models from other classes may work as well but are not officially supported. The model should include the interaction of interest.- pred
A categorical predictor variable that will appear on the x-axis. Note that it is evaluated using
rlang, so programmers can use the!!syntax to pass variables instead of the verbatim names.- modx
A categorical moderator variable.
- mod2
For three-way interactions, the second categorical moderator.
- data
Optional, default is NULL. You may provide the data used to fit the model. This can be a better way to get mean values for centering and can be crucial for models with variable transformations in the formula (e.g.,
log(x)) or polynomial terms (e.g.,poly(x, 2)). You will see a warning if the function detects problems that would likely be solved by providing the data with this argument and the function will attempt to retrieve the original data from the global environment.- geom
What type of plot should this be? There are several options here since the best way to visualize categorical interactions varies by context. Here are the options:
"point": The default. Simply plot the point estimates. You may want to usepoint.shape = TRUEwith this and you should also considerinterval = TRUEto visualize uncertainty."line": This connects observations across levels of thepredvariable with a line. This is a good option when thepredvariable is ordinal (ordered). You may still considerpoint.shape = TRUEandinterval = TRUEis still a good idea."bar": A bar chart. Some call this a "dynamite plot." Many applied researchers advise against this type of plot because it does not represent the distribution of the observed data or the uncertainty of the predictions very well. It is best to at least use theinterval = TRUEargument with this geom.
- pred.values
Which values of the predictor should be included in the plot? By default, all levels are included.
- modx.values
For which values of the moderator should lines be plotted? There are two basic options:
A vector of values (e.g.,
c(1, 2, 3))A single argument asking to calculate a set of values. See details below.
Default is
NULL. IfNULL(ormean-plus-minus), then the customary +/- 1 standard deviation from the mean as well as the mean itself are used for continuous moderators. If"plus-minus", plots lines when the moderator is at +/- 1 standard deviation without the mean. You may also choose"terciles"to split the data into equally-sized groups and choose the point at the mean of each of those groups.If the moderator is a factor variable and
modx.valuesisNULL, each level of the factor is included. You may specify any subset of the factor levels (e.g.,c("Level 1", "Level 3")) as long as there is more than 1. The levels will be plotted in the order you provide them, so this can be used to reorder levels as well.- mod2.values
For which values of the second moderator should the plot be facetted by? That is, there will be a separate plot for each level of this moderator. Defaults are the same as
modx.values.- interval
Logical. If
TRUE, plots confidence/prediction intervals.- plot.points
Logical. If
TRUE, plots the actual data points as a scatterplot on top of the interaction lines. Note that ifgeom = "bar", this will cause the bars to become transparent so you can see the points.- point.shape
For plotted points—either of observed data or predicted values with the "point" or "line" geoms—should the shape of the points vary by the values of the factor? This is especially useful if you aim to be black and white printing- or colorblind-friendly.
- vary.lty
Should the resulting plot have different shapes for each line in addition to colors? Defaults to
TRUE.- centered
A vector of quoted variable names that are to be mean-centered. If
"all", all non-focal predictors are centered. You may instead pass a character vector of variables to center. User can also use "none" to base all predictions on variables set at 0. The response variable,pred,modx, andmod2variables are never centered.- int.type
Type of interval to plot. Options are "confidence" or "prediction". Default is confidence interval.
- int.width
How large should the interval be, relative to the standard error? The default, .95, corresponds to roughly 1.96 standard errors and a .05 alpha level for values outside the range. In other words, for a confidence interval, .95 is analogous to a 95% confidence interval.
- line.thickness
How thick should the plotted lines be? Default is 1.
- point.size
What size should be used for observed data when
plot.pointsis TRUE? Default is 1.5.- pred.point.size
If TRUE and
geomis"point"or"line", sets the size of the predicted points. Default is 3.5. Note the distinction frompoint.size, which refers to the observed data points.- jitter
How much should
plot.pointsobserved values be "jittered" viaggplot2::position_jitter()? When there are many points near each other, jittering moves them a small amount to keep them from totally overlapping. In some cases, though, it can add confusion since it may make points appear to be outside the boundaries of observed values or cause other visual issues. Default is 0.1, but increase as needed if your points are overlapping too much or set to 0 for no jitter. If the argument is a vector with two values, then the first is assumed to be the jitter for width and the second for the height.- geom.alpha
What should the alpha aesthetic be for the plotted lines/bars? Default is NULL, which means it is set depending on the value of
geomandplot.points.- dodge.width
What should the
widthargument toggplot2::position_dodge()be? Default is NULL, which means it is set depending on the value ofgeom.- errorbar.width
How wide should the error bars be? Default is NULL, meaning it is set depending on the value
geom. Ignored ifintervalis FALSE.- interval.geom
For categorical by categorical interactions. One of "errorbar" or "linerange". If the former,
ggplot2::geom_errorbar()is used. If the latter,ggplot2::geom_linerange()is used.- outcome.scale
For nonlinear models (i.e., GLMs), should the outcome variable be plotted on the link scale (e.g., log odds for logit models) or the original scale (e.g., predicted probabilities for logit models)? The default is
"response", which is the original scale. For the link scale, which will show straight lines rather than curves, use"link".- robust
Should robust standard errors be used to find confidence intervals for supported models? Default is FALSE, but you should specify the type of sandwich standard errors if you'd like to use them (i.e.,
"HC0","HC1", and so on). IfTRUE, defaults to"HC3"standard errors.- cluster
For clustered standard errors, provide the column name of the cluster variable in the input data frame (as a string). Alternately, provide a vector of clusters.
- vcov
Optional. You may supply the variance-covariance matrix of the coefficients yourself. This is useful if you are using some method for robust standard error calculation not supported by the sandwich package.
- pred.labels
A character vector of equal length to the number of factor levels of the predictor (or number specified in
predvals). IfNULL, the default, the factor labels are used.- modx.labels
A character vector of labels for each level of the moderator values, provided in the same order as the
modx.valuesargument. IfNULL, the values themselves are used as labels unlessmodx,valuesis alsoNULL. In that case, "+1 SD" and "-1 SD" are used.- mod2.labels
A character vector of labels for each level of the 2nd moderator values, provided in the same order as the
mod2.valuesargument. IfNULL, the values themselves are used as labels unlessmod2.valuesis alsoNULL. In that case, "+1 SD" and "-1 SD" are used.- set.offset
For models with an offset (e.g., Poisson models), sets an offset for the predicted values. All predicted values will have the same offset. By default, this is set to 1, which makes the predicted values a proportion. See details for more about offset support.
- x.label
A character object specifying the desired x-axis label. If
NULL, the variable name is used.- y.label
A character object specifying the desired x-axis label. If
NULL, the variable name is used.- main.title
A character object that will be used as an overall title for the plot. If
NULL, no main title is used.- legend.main
A character object that will be used as the title that appears above the legend. If
NULL, the name of the moderating variable is used.- colors
Any palette argument accepted by
jtools::get_colors(). Default is "CUD Bright" for factor moderators and "blue" for continuous moderators. You may also simply supply a vector of colors accepted byggplot2and of equal length to the number of moderator levels.- partial.residuals
Instead of plotting the observed data, you may plot the partial residuals (controlling for the effects of variables besides
pred).- point.alpha
What should the
alphaaesthetic for plotted points of observed data be? Default is 0.6, and it can range from 0 (transparent) to 1 (opaque).- color.class
Deprecated. Now known as
colors.- at
If you want to manually set the values of other variables in the model, do so by providing a named list where the names are the variables and the list values are vectors of the values. This can be useful especially when you are exploring interactions or other conditional predictions.
- ...
extra arguments passed to
make_predictions
Value
The functions returns a ggplot object, which can be treated
like a user-created plot and expanded upon as such.
Details
This function provides a means for plotting conditional effects
for the purpose of exploring interactions in the context of regression.
You must have the
package ggplot2 installed to benefit from these plotting functions.
The function is designed for two and three-way interactions. For
additional terms, the
effects package may be better suited to the task.
This function supports nonlinear and generalized linear models and by
default will plot them on
their original scale (outcome.scale = "response").
While mixed effects models from lme4 are supported, only the fixed
effects are plotted. lme4 does not provide confidence intervals,
so they are not supported with this function either.
Note: to use transformed predictors, e.g., log(variable),
provide only the variable name to pred, modx, or mod2 and supply
the original data separately to the data argument.
Info about offsets:
Offsets are partially supported by this function with important
limitations. First of all, only a single offset per model is supported.
Second, it is best in general to specify offsets with the offset argument
of the model fitting function rather than in the formula. You are much
more likely to have success if you provide the data used to fit the model
with the data argument.
Examples
library(ggplot2)
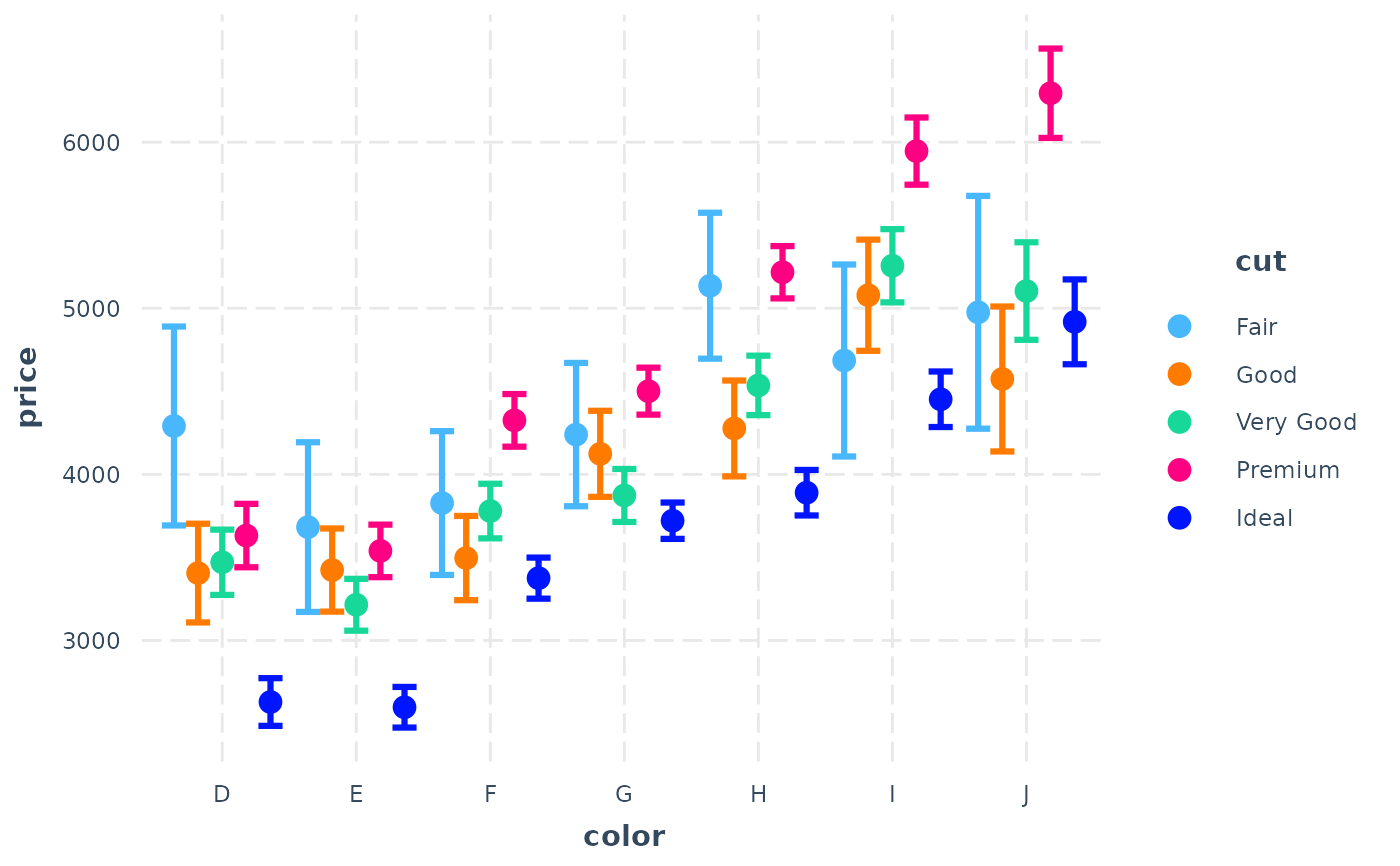
fit <- lm(price ~ cut * color, data = diamonds)
cat_plot(fit, pred = color, modx = cut, interval = TRUE)
 # 3-way interaction
## Will first create a couple dichotomous factors to ensure full rank
mpg2 <- mpg
mpg2$auto <- "auto"
mpg2$auto[mpg2$trans %in% c("manual(m5)", "manual(m6)")] <- "manual"
mpg2$auto <- factor(mpg2$auto)
mpg2$fwd <- "2wd"
mpg2$fwd[mpg2$drv == "4"] <- "4wd"
mpg2$fwd <- factor(mpg2$fwd)
## Drop the two cars with 5 cylinders (rest are 4, 6, or 8)
mpg2 <- mpg2[mpg2$cyl != "5",]
mpg2$cyl <- factor(mpg2$cyl)
## Fit the model
fit3 <- lm(cty ~ cyl * fwd * auto, data = mpg2)
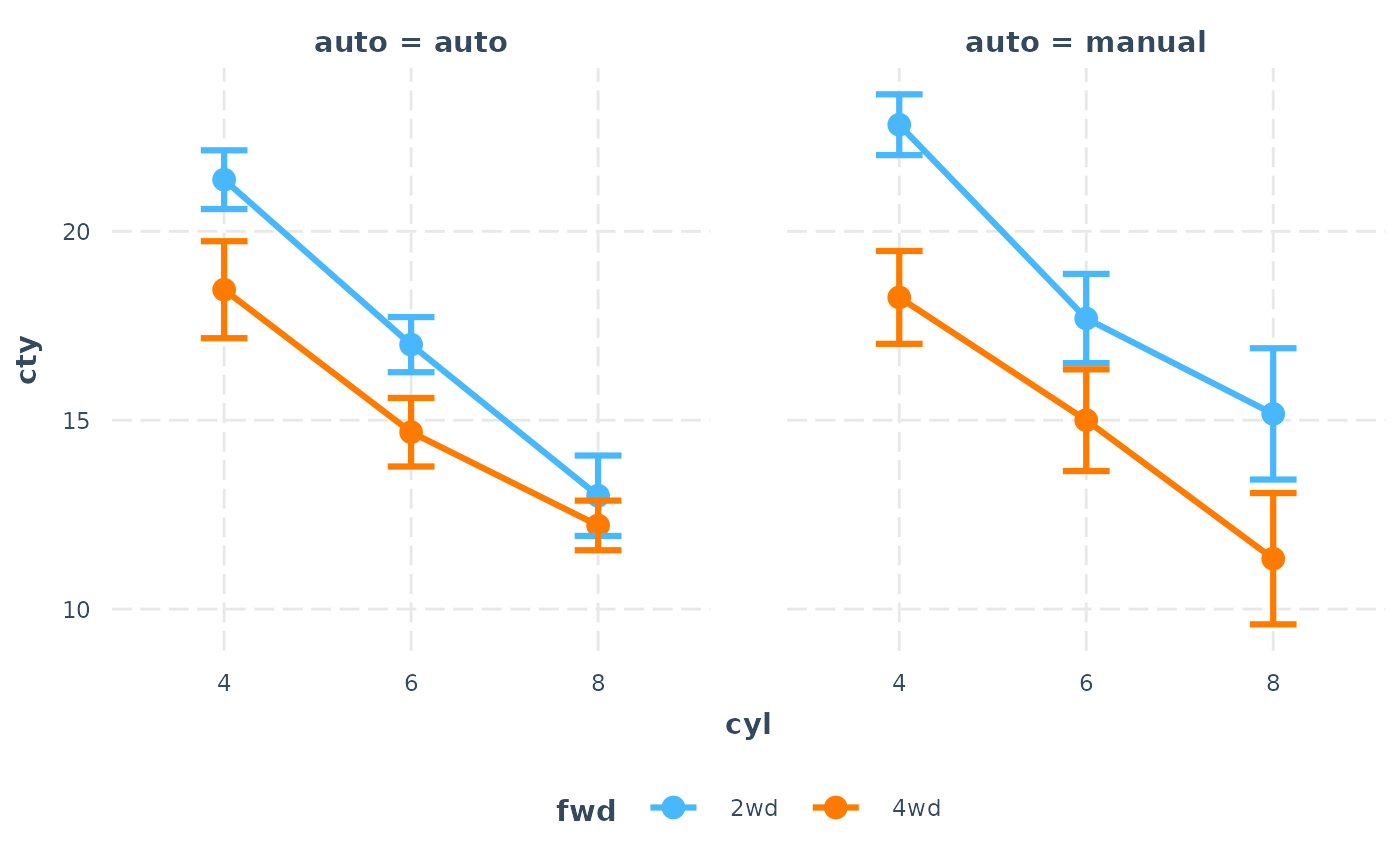
# The line geom looks good for an ordered factor predictor
cat_plot(fit3, pred = cyl, modx = fwd, mod2 = auto, geom = "line",
interval = TRUE)
# 3-way interaction
## Will first create a couple dichotomous factors to ensure full rank
mpg2 <- mpg
mpg2$auto <- "auto"
mpg2$auto[mpg2$trans %in% c("manual(m5)", "manual(m6)")] <- "manual"
mpg2$auto <- factor(mpg2$auto)
mpg2$fwd <- "2wd"
mpg2$fwd[mpg2$drv == "4"] <- "4wd"
mpg2$fwd <- factor(mpg2$fwd)
## Drop the two cars with 5 cylinders (rest are 4, 6, or 8)
mpg2 <- mpg2[mpg2$cyl != "5",]
mpg2$cyl <- factor(mpg2$cyl)
## Fit the model
fit3 <- lm(cty ~ cyl * fwd * auto, data = mpg2)
# The line geom looks good for an ordered factor predictor
cat_plot(fit3, pred = cyl, modx = fwd, mod2 = auto, geom = "line",
interval = TRUE)